seq_to_satn#
Converts values of invasion sequence into a saturation map
import numpy as np
import porespy as ps
import matplotlib.pyplot as plt
from edt import edt
ps.visualization.set_mpl_style()
Generate an image containing invasion sizes using the porosimetry function:
np.random.seed(0)
im = ps.generators.blobs([200, 200], porosity=0.5)
inv = ps.filters.porosimetry(im)
Then convert the sizes to sequence values:
seq = ps.filters.size_to_seq(inv, im=im)
seq#
satn = ps.filters.seq_to_satn(seq=seq, im=im)
fig, ax = plt.subplots(1, 2, figsize=[8, 4])
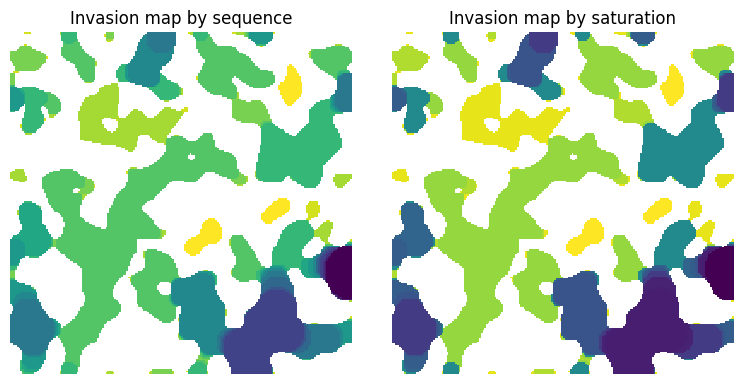
ax[0].imshow(seq / im, origin="lower", interpolation="none")
ax[0].set_title("Invasion map by sequence")
ax[0].axis(False)
ax[1].imshow(satn / im, origin="lower", interpolation="none")
ax[1].set_title("Invasion map by saturation")
ax[1].axis(False);
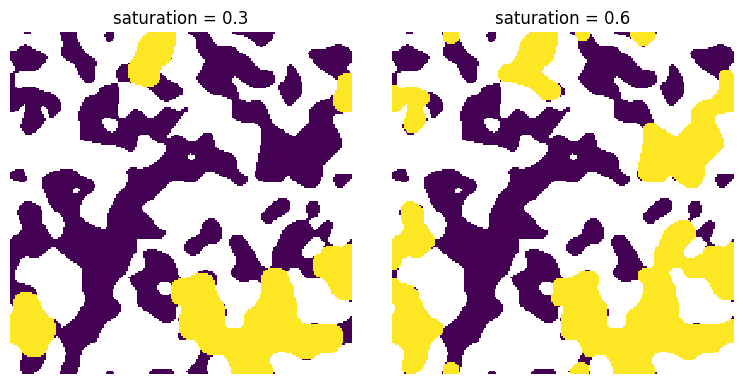
The saturation map makes it very easy to obtain a desired fluid configuration just by applying a threhold:
fig, ax = plt.subplots(1, 2, figsize=[8, 4])
s = 0.3
ax[0].imshow((satn < s) * (satn > 0) / im, origin="lower", interpolation="none")
ax[0].set_title(f"saturation = {s}")
ax[0].axis(False)
s = 0.6
ax[1].imshow((satn < s) * (satn > 0) / im, origin="lower", interpolation="none")
ax[1].set_title(f"saturation = {s}")
ax[1].axis(False);