label_phases#
Version 2 of PoreSpy included the ability to perform network extractions on images that contain multiple phases, as outlined by Khan et al. The regions_to_network function includes the ability to label each pore with the phase to which it belongs, but does nothing else. The label_phases function then analyzes the network output by regions_to_network to create the labels that can be used within OpenPNM.
import numpy as np
import porespy as ps
np.random.seed(13)
im = ps.generators.overlapping_spheres([100, 100], r=7, porosity=0.7)
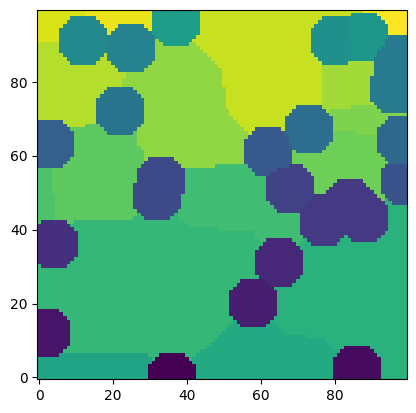
snow = ps.filters.snow_partitioning_n(im=im.astype(int) + 1)
ps.imshow(snow.regions, origin="lower", interpolation="none");
network#
The dictionary returned from the regions_to_network function must be supplied:
net = ps.networks.regions_to_network(regions=snow.regions, phases=snow.im)
net = ps.networks.label_phases(network=net)
for item in net.keys():
print(item)
throat.conns
pore.coords
pore.all
throat.all
pore.region_label
pore.phase
throat.phases
pore.region_volume
pore.equivalent_diameter
pore.local_peak
pore.global_peak
pore.geometric_centroid
throat.global_peak
pore.inscribed_diameter
pore.extended_diameter
throat.inscribed_diameter
throat.total_length
throat.direct_length
throat.perimeter
pore.volume
pore.surface_area
throat.cross_sectional_area
throat.equivalent_diameter
pore.void
throat.void_void
throat.void_solid
pore.solid
throat.solid_void
throat.solid_solid
In the above print-out we can see that several labels have been added to the list, such as 'throat.void_void' which is True for all throats which connect a void pore to another void pore, and so forth.
alias#
We can override the default names of 'solid' and 'void' by providing a dict which maps the phase number to our desired name as follows:
net = ps.networks.regions_to_network(regions=snow.regions, phases=snow.im)
net = ps.networks.label_phases(network=net, alias={1: "void", 2: "grain"})
for item in net.keys():
print(item)
throat.conns
pore.coords
pore.all
throat.all
pore.region_label
pore.phase
throat.phases
pore.region_volume
pore.equivalent_diameter
pore.local_peak
pore.global_peak
pore.geometric_centroid
throat.global_peak
pore.inscribed_diameter
pore.extended_diameter
throat.inscribed_diameter
throat.total_length
throat.direct_length
throat.perimeter
pore.volume
pore.surface_area
throat.cross_sectional_area
throat.equivalent_diameter
pore.void
throat.void_void
throat.void_grain
pore.grain
throat.grain_void
throat.grain_grain
Now we can see that 'solid' and 'void' have been replaced by 'void' and 'grain'.
